Research Library
The top resource for free research, white papers, reports, case studies, magazines, and eBooks.
- Agriculture
- Automotive
- Career
- Construction
- Education
- Engineering
- Finance
- Food and Beverage
- Government
- Healthcare and Medical
- Human Resources
- Information Technology
- Data Infrastructure
- Data Tools
- Desktops, Laptops and OS
- Chip Sets
- Collaboration Tools
- Desktop Systems - PCs
- Email Client
- Embedded Systems
- Hardware and Periferals
- Laptops
- Linux - Open Source
- Mac OS
- Memory Components
- Mobile Devices
- Presentation Software
- Processors
- Spreadsheets
- Thin Clients
- Upgrades and Migration
- Windows 7
- Windows Vista
- Windows XP
- Word Processing
- Workstations
- Enterprise Applications
- IT Infrastructure
- IT Management
- Networking and Communications
- Bluetooth
- DSL
- GPS
- GSM
- Industry Standard Protocols
- LAN - WAN
- Management
- Mobile - Wireless Communications
- Network
- Network Administration
- Network Design
- Network Disaster Recovery
- Network Interface Cards
- Network Operating Systems
- PBX
- RFID
- Scalability
- TCP - IP
- Telecom Hardware
- Telecom Regulation
- Telecom Services
- Telephony Architecture
- Unified Communications
- VPNs
- VoIP - IP Telephony
- Voice Mail
- WAP
- Wi-Fi (802.11)
- WiMAX (802.16)
- Wide Area Networks (WAN)
- Wireless Internet
- Wireless LAN
- Security
- Servers and Server OS
- Software and Web Development
- .Net Framework
- ASPs
- Application Development
- Application Servers
- Collaboration
- Component-Based
- Content Management
- E-Commerce - E-Business
- Enterprise Applications
- HTML
- IM
- IP Technologies
- Integration
- Internet
- Intranet
- J2EE
- Java
- Middleware
- Open Source
- Programming Languages
- Quality Assurance
- SAAS
- Service-Oriented Architecture (SOA)
- Software Engineering
- Software and Development
- Web Design
- Web Design and Development
- Web Development and Technology
- XML
- Storage
- Life Sciences
- Management
- Manufacturing
- Marketing
- Meetings and Travel
- Multimedia
- Operations
- Retail
- Sales
- Trade/Professional Services
- Utility and Energy
- View All Topics
- Featured eBooks
- Trending Resources
- New Resources
- Promote Your Content
- Partnership Opportunities
- Get RSS Updates
- About TradePub.com
- FAQ
- Contact Us
Share Your Content with Us
on TradePub.com for readers like you. LEARN MORE
Request Your Free White Paper Now:
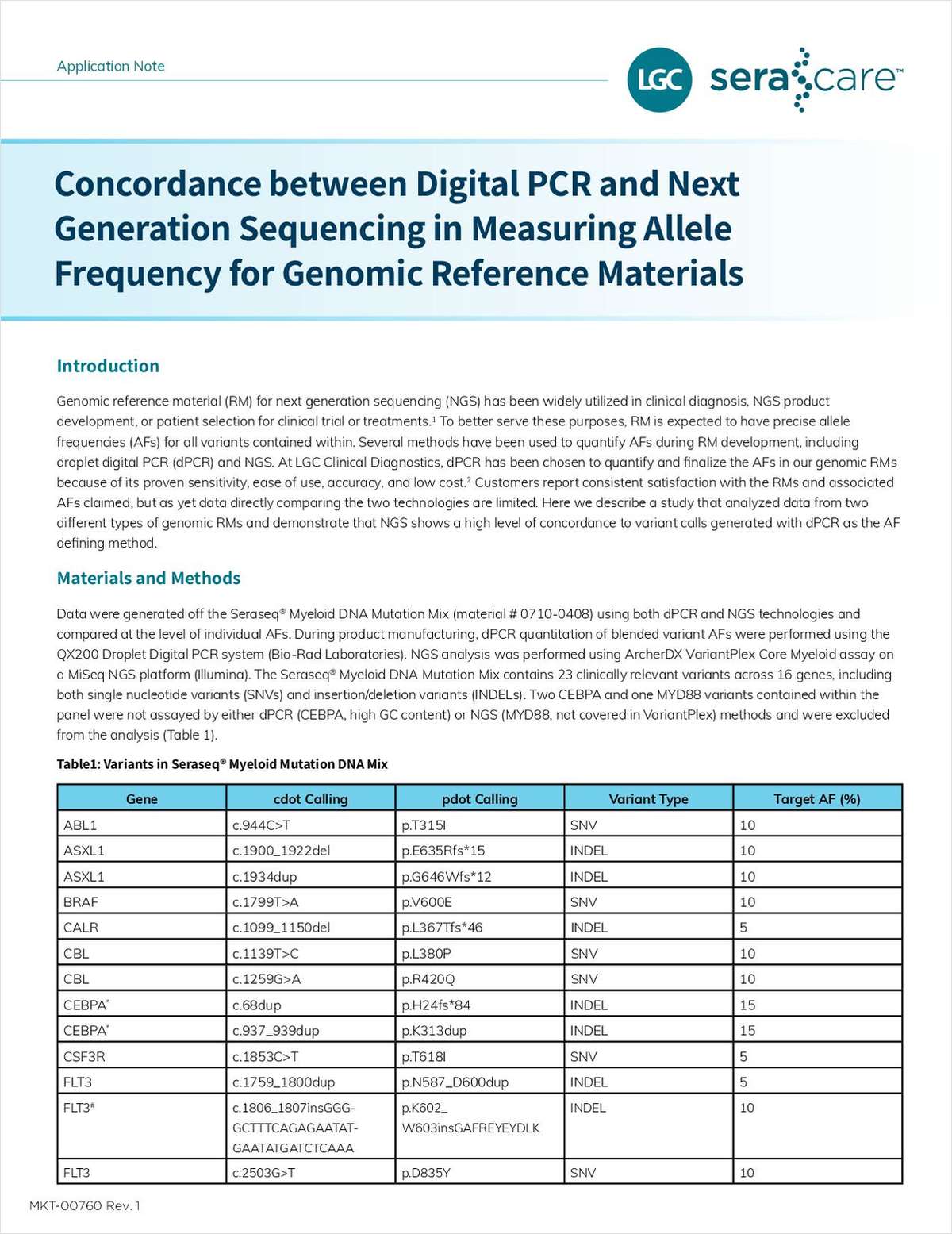
"Concordance Between Digital PCR and Next-Generation Sequencing in Measuring Allele Frequency for Genomic Reference Materials"
Genomic reference material (RM) for next-generation sequencing (NGS) has been widely utilized in clinical diagnosis, NGS product development, and patient selection for clinical trials or treatments. To better serve these purposes, RM is expected to have precise allele frequencies (AFs) for all variants contained within.
Several methods have been used to quantify AFs during RM development, including droplet digital PCR (dPCR) and NGS. At LGC Clinical Diagnostics, dPCR has been chosen to quantify and finalize the AFs in our genomic RMs because of its proven sensitivity, ease of use, accuracy, and low cost. Customers report consistent satisfaction with the RMs and associated AFs claimed, but as yet, data directly comparing the two technologies are limited.
This white paper from LGC SeraCare describes a study that analyzed data from two different types of genomic reference materials and demonstrates that next-generation sequencing shows a high level of concordance to variant calls generated with droplet digital PCR as the precise allele frequency-defining method.
Offered Free by: LGC SeraCare Life Sciences
See All Resources from: LGC SeraCare Life Sciences